Performance¶
This page is generated by a Jupyter notebook which can be opened and run in Binder or Google Colab by clicking on the above badges. To run it in Google Colab, you need to install PyMinimax in Colab first:
[ ]:
pip install pyminimax
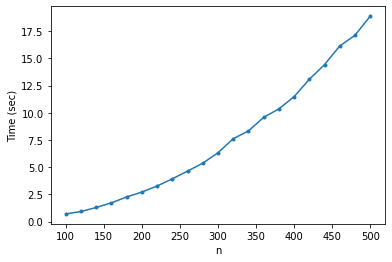
The most computationally intensive function in PyMinimax is pyminimax.minimax. Although implemented as an efficient nearest-neighbor chain algorithm, PyMinimax is a pure Python implementation at least for now, so it is relatively slow. Below is a plot of the running time on Binder against size of the dataset.
When \(n=3000\), pyminimax.minimax takes a little more than 10 minutes on Binder to compute the linkage matrix. For now PyMinimax is only suitable for small to medium datasets.
[1]:
import time
import numpy as np
import matplotlib.pyplot as plt
from pandas import DataFrame
from pyminimax import minimax
from scipy.spatial.distance import pdist
np.random.seed(0)
n_pts = np.arange(100, 501, 20)
elapsed_time = []
def test_run(n):
X = np.random.rand(n, 2)
Z = minimax(pdist(X))
for n in n_pts:
start = time.process_time()
test_run(n)
end = time.process_time()
elapsed_time.append(end - start)
df = DataFrame({'n': n_pts, 'time': elapsed_time}).set_index('n')
ax = df.plot(style='.-', legend=None);
ax.set(ylabel='Time (sec)')
plt.show()